In the soon-to-be-released sDNA3, it will no longer necessary to repair split links. This looks like a departure from how sDNA used to work, so I’d better explain it.
First, let’s clarify what we’re talking about. A split link is where a single network link is represented by two or more polylines in the data, rather than a single polyline as is customary. Here’s an example, in which I’ve placed circles over polyline ends to make them visible. The polylines shown in red form a split link. (Of course they look like a single line – we only know it’s a split link because of the node halfway along).
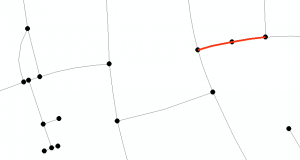
Base data (c) Openstreetmap contributors
The point where the two red links join is often called a pseudonode in the technical literature, so we can also say that split links are the lines joined by pseudonodes.
Two of our aims in making the original sDNA tool were (i) to exploit measures of link density to add information to our models, and (ii) to be compatible with data in the format everybody uses, that is to say, link-node. For this reason it seemed like a good idea to insist on using whole links as the unit of analysis, so we specified that sDNA must be fed networks without split links, and provided a tool for repairing them.
It turns out that was a bad idea, as it doesn’t offer our users enough flexibility. In our own work, and also working with our Impact Accelerator partners, we have found several reasons why you might want to leave split links as they are:
- Topology checking. Accidental intersections between lines in network data cause huge problems for network analysis, as they imply a network model which is wrong. Unfortunately they are also hard to spot, because on most maps they look just like lines with coincident endpoints. For this reason, many data providers will split ALL links where they cross, even where they don’t join (e.g. bridges and tunnels – to which grade separation data is added to show what is going on). Working this way has the advantage that any intersections between lines are definitely an error, so errors are easier to find with a topology checker.
- Representing sub-links with different characteristics. For example, a one way system, toll charge, major origin or destination, etc, that applies only to part of a link.
- Obtaining output at sub-link level. If many origins and destinations fall on a link, traffic flow will vary along the length of the link, and you may wish to model this rather than treat the link as a single entity. Or, you may wish to measure accessibility at sub-link level.
- Sub-link accuracy for routing algorithms. In some cases, somebody’s starting position on a link will affect the way they choose to leave it. For most models this does not matter, but if it does in yours, you may want to divide links into small chunks to represent it.
- Testing different network design options. Ultimately, sDNA exists to help designers plan better networks, so testing out different alternatives is part of the job. While you could prepare an entire new network for each option you want to test, it’s usually easier to add and remove a small set of links by switching them on or off. Another new feature of sDNA3 is that it allows you to do this. However, switching off a link will usually have the effect of creating pseudonodes, and hence split links, at its endpoints.
So, here is how split links are handled in sDNA3:
- sDNA still uses link-node format.
- Links can, however, be composed of one of more polylines. At your choice you can either use one, or more than one polyline to represent each link. If you choose to use only one polyline per link, you are using sDNA in the way we used to recommend, but you have other options now.
- Each polyline is treated as a single entity for routing decisions, that is to say, all traffic between each pair of polylines will go the same way (unless you use random hybrid metrics and oversampling).
- Three forms of weighting are offered:
- Polyline weighting: each polyline receives a weight of 1 (unweighted), or a custom weight which applies directly to the polyline.
- Length weighting: each polyline receives a weight equal to its length (unweighted), or if using custom weights, these are taken to represent weight per unit length.
- Link weighting (the default): each link receives a weight of 1 (unweighted), or if using custom weights, they are taken to represent weight per link. In the case of split links, the weight is divided over the polylines that form the link, proportional to their length. Custom weights for parts of split links are scaled down according to the fraction of the link they represent.
Finally, sDNA3 outputs a new measurement called LFrac or Link Fraction. This shows you what proportion of a link each polyline in your model represents.
Link weighting sounds complicated, but really it’s the same thing we were doing from the beginning. If all split links are repaired, then polyline weighting is the same as link weighting; which is why in sDNA versions 1 and 2 we achieved link weighting by using polyline weighting and providing you with a tool to repair the split links.
There are, of course, reasons to use the other types of weighting on occasion. Polyline weighting is useful if you have address point data which you have attached to the network, as it preserves the exact value of each attached weight. Length weighting is useful for analyses where you think network length is more representative of urban density than link counts. Both of the latter can also be used along with land use data to provide custom weights per link, or per unit length, for different types of land use.
But go ahead and use split links: leaving them in there won’t affect the way the analysis is weighted by default. If you use a lot of split links (and I mean a lot – like splitting every link into multiple parts) it will slow down sDNA, but if you only use a few here and there – say for bridges, design options, one way systems, etc – you won’t notice the difference in speed. If you want higher accuracy only on a small part of your model, you can also split links in that part only to keep compute times down.
